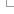Chromosomes, scaffolds and contigs
Genome assemblies are hierarchical. The shortest assembly components are contigs, which are sequences taken from individuals. Contigs are assembled into longer scaffolds, and scaffolds are assembled into chromosomes if there is sufficient mapping information. Many genome assemblies have only been assembled to the scaffold level.
Scaffolds are classified in three ways:
- Placed scaffolds: the scaffolds have been placed within a chromosome.
- Unlocalised scaffolds: although the chromosome within which the scaffold occurs is known, the scaffold's position or orientation is not known.
- Unplaced scaffolds: it is not known which chromosome the scaffold belongs to.
The relationship between contigs, scaffolds and chromosomes is defined in AGP files. These files describe how assembled sequences (eg chromosomes) are compiled from their components (eg scaffolds). In Ensembl, we import contig-level DNA sequence into our core databases. We also import the AGP files for contig-to-scaffold, contig-to-chromosome, and scaffold-to-chromosome mappings. This allows us to generate scaffold and chromosome sequence on the fly by stitching the contigs sequences together as specified by the AGP files.
Toplevel
For each genome assembly, we define the set of toplevel sequences. These are sequence regions in the genome assembly that are not a component of another sequence region. For example, when a genome is assembled into chromosomes, toplevel sequences will be chromosomes and any unlocalised or unplaced scaffolds. If a genome has only been assembled into scaffolds, then toplevel sequences are the full set of unlocalised and unplaced scaffolds.